Biomedical Network Science Lab
The Biomedical Network Science (BIONETS) Lab investigates molecular mechanisms, using techniques from network science, (graph-based) AI and combinatorial optimization. We develop algorithms, AI models, and software tools to mine omics data for such mechanisms, with the aim to better understand cellular pathways and, ultimately, pave the way for targeted and causally effective treatments of complex diseases. We also develop privacy-preserving decentralized biomedical AI solutions, which enable cross-institutional studies on sensitive data. Finally, we are interested in meta-scientific questions such as reproducibility and the impact of data bias on biomedical AI systems.

News
Welcome Dominik

On April 1st, Dominik Pysch joined the BIONETS Lab as a new PhD candidate. He will work on graph-based modelling of tissue inflammation in the context of FAU’s and UKER’s new CRC CASCAID.
Welcome Judith

We are very happy that, on March 1st, Judith Bernett joined the BIONETS Lab as a new research associate. Judith will work on ML models for context-specific filtering of protein-protein interaction networks in the context of the CoBiNet project (https://cobinet.ai).
Productive discussions at the DyHealthNet workshop in Erlangen

More than 30 researchers and project partners gathered in Erlangen for the DyHealthNet workshop on March 19-20. Within these two days, several interesting presentations and discussions from professionals in the field of population cohorts took place, including researchers from the SHIP and FinnGen study, and the Pan-Canadian Genome Library. We also presented our experiences from…
AIBeez

We are very happy to announce the birth of AIBeez – “Association for the Promotion of Artificial Intelligence in Health and Engineering Sciences in Erlangen e.V.” New members who want to support AI research for biomedicine in Erlangen are very welcome!
Paper published in Genome Biology
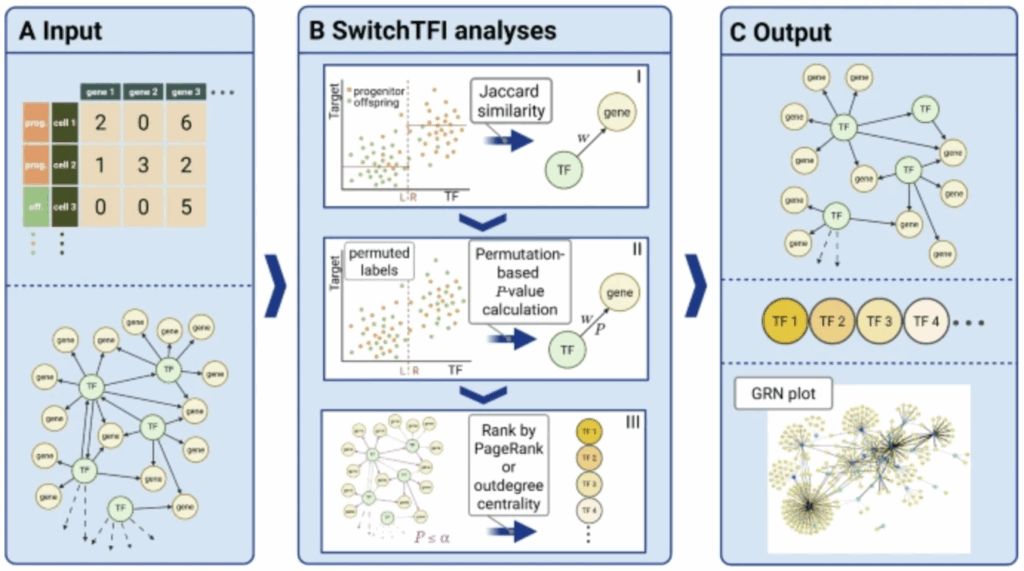
In our paper “SwitchTFI: identifying transcription factors driving cell differentiation”, we present an algorithm to infer regulatory mechanisms that drive the transition from progenitor to offspring cells from scRNA-seq data.
DFG funds CASCAID

We are very excited to announce that the DFG funds the new CRC “Cellular and Systems Control of Autoimmune Disease” (CASCAID). Together with the lab of Stefan Uderhardt, we will develop graph-based models of histological healing that quantify the target of successful treatment of autoimmune diseases. Moreover, we will provide CASCAID-wide bioinformatics and data analysis…
Paper published in GigaScience

In our paper “NApy: Efficient Statistics in Python for Large-Scale Heterogeneous Data with Enhanced Support for Missing Data”, we present a Python package with Numba and OpenMP-powered C++ backends for fast and memory-efficient statistical testing in the presence of missing data.
Paper published in ISMB 2025 proceedings
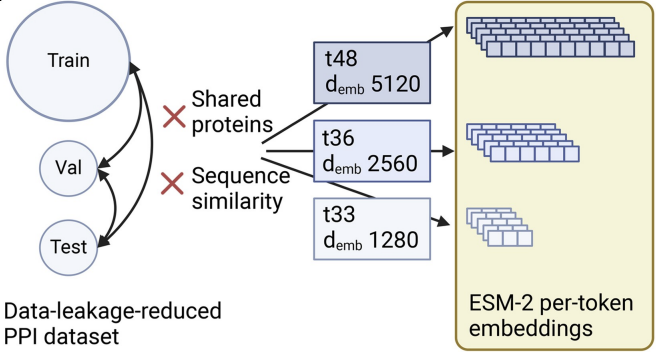
In our paper “Deep learning models for unbiased sequence-based PPI prediction plateau at an accuracy of 0.65” we show that usage of ESM-2 embeddings boosts performance in out-of-distribution PPI prediction to around 0.65 independently of model architecture.
Paper published in Bioinformatics
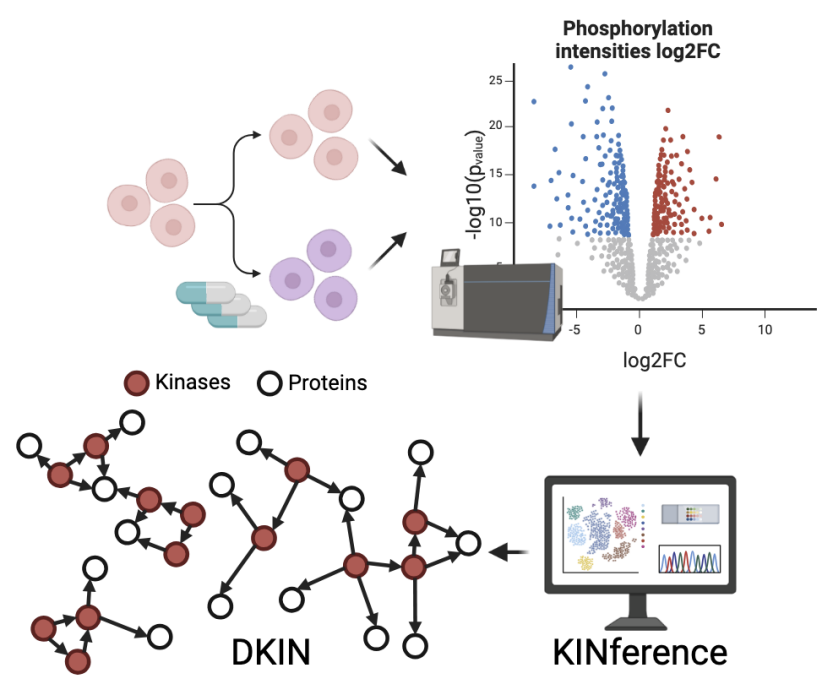
In our paper “Inference of differential kinase interaction networks with KINference”, we present the R tool KINference, which can be used to identify kinase-substrate links that are differentially active between two conditions.
